Current Projects
Computational modeling of interactions between bacteria and immune cells in tumors
With the development of synthetic biology, tumor‑targeting bacterial therapy is becoming more plausible as an alternative treatment for conventional cancer therapy. Scientists noticed that some immune cells have different responses after bacteria injection. Understanding the mechanism of bacteria-immune cell interactions is crucial for enhancing the effectiveness of bacterial therapy.
To address this question, I am developing an individual-based model framework to simulate the interactions between bacteria and immune cells in tumors and comparing the simulation results with experimental data. I am also attempting to use reaction-diffusion models to simulate these interactions from a different point of view.
Tracking cells in PDMS microfluidic chips in phase-contrast images

This project is a collaboration with Fu lab and the project background is the same as the above project.
The main work I have done in this project is tracking cells from videos. I wrote the code independently for this project. Check code here.
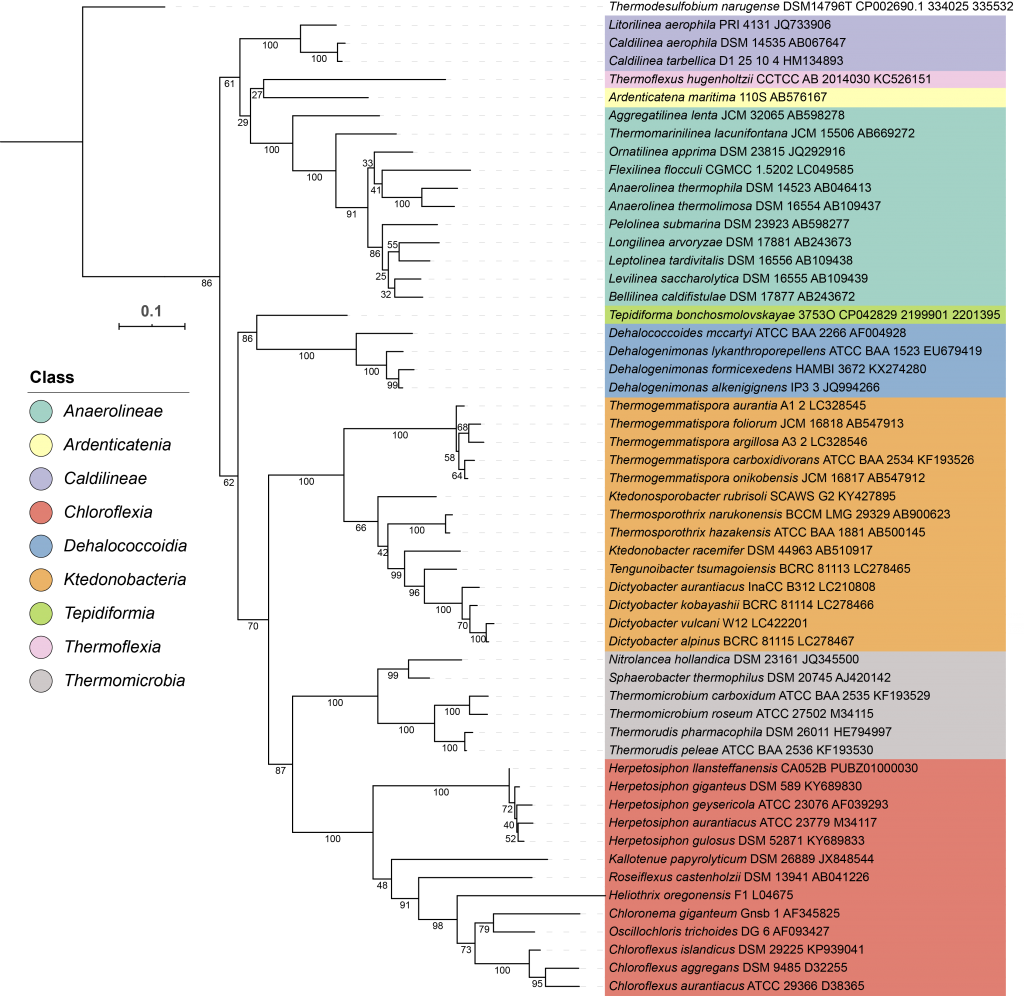
Global distribution and genomic inference of the metabolism of the classTepidiformia in the phylum Chloroflexi
Bacteria in the phylum Chloroflexi are widespread in free-living microbial communities, and it is famous for one special carbon fixation pathway that exclusively exists in this phylum, called 3-hydroxypropionate bicycle. The class Tepidiformia is a newly discovered member in the phylum Chloroflexi in 2019. I used the metagenome-assembled genomes from the hot springs derived metagenomic data of my lab to search for the existence of Tepidiformia. I analyzed the global distribution and metabolic potentials of this class. The paper is in preparation.
Update on the classification of higher ranks in the phylum Actinobacteria
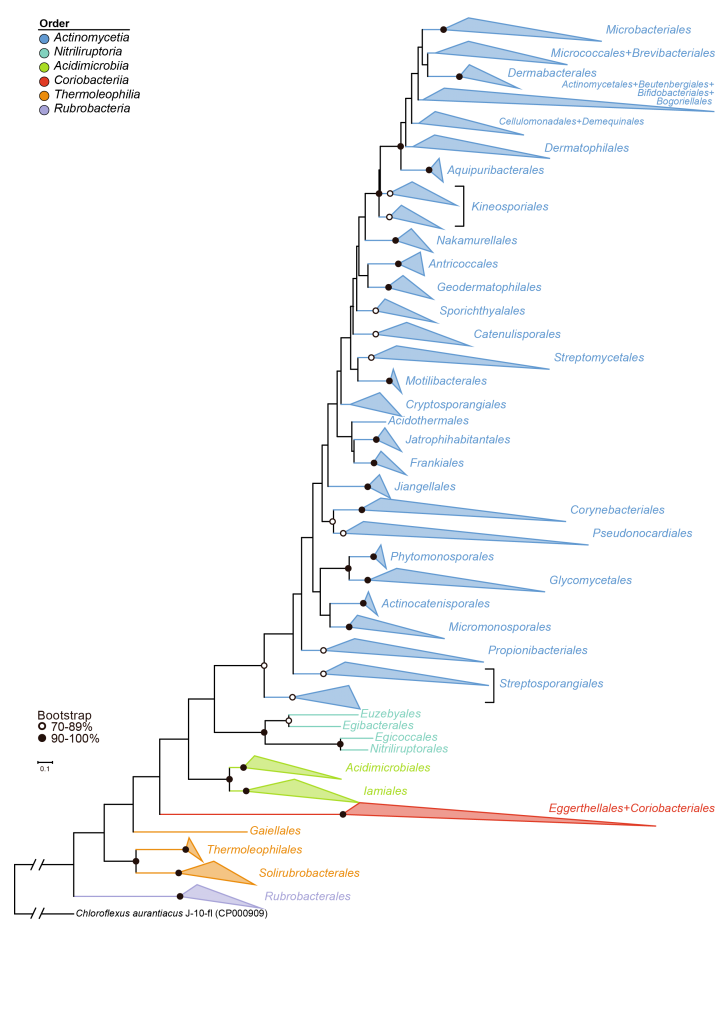
The phylum Actinobacteria is one of the largest lineages in the domain Bacteria. It has accumulated many new species with 16S rRNA gene sequences and genomic data during the past decade, while the phylogenetic relationships of this phylum may need to be reconstructed. So we proposed a rearrangement of the higher taxonomic ranks of the phylum Actinobacteria based on the 16S rRNA gene sequences and genome sequences. See more
Previous Projects
This part is under construction.